RFP Lab

Purpose
Make RFP from jelly fish in bacteria
Learn about steps of genetic engineering
Materials and Procedure
2a- materials and procedure can be found in Amgen manual part 2a
4a- materials and procedure can be found in Amgen manual part 4a
5a- materials and procedure can be found in Amgen manual part 5a
6a- materials and procedure can be found in Amgen manual part 6a
Experimental Overview
Part 2a: Use restriction digest to verify plasmid. We cut the plasmid with Hindlll and BamH1, to have the ability to get a portion of the bacteria gene.
Part 4a: Verification of plasmid digest by electrophoresis. We used gel electrophoresis in which a plasmid is ran along a DNA ladder. This ladder has bands that go hand-in-hand with other spaces or lane to determine the molecules stature. We know the size of the RFP gene so we knew how well the digest had ran. The digest ran perfectly.
Part 5a: Transformation of bacteria with recombinant plasmid. We cut plasmids with enzymes specifically restriction enzymes and to put together the gene of interest we used a substance called ligase.
Part 6a: We incubated the RFP gene or kept it warm and we separated the RFP from everything else. Next we used hydrophilic beads to collect the RFP and finally we transported the RFP and sort of made it mad by striking it against a pipet rack.
Data
2a- Why is it important that the same enzyme or
enzymes be used to cut both the plasmid and the insulin gene from the
human DNA?
If we cut the plasmid and the insulin gene from the DNA so the ends of them connect and fit together like puzzle pieces.
Why does using two different enzymes to cut the
plasmid prevent the plasmid from reforming a circle without the
inserted gene?
Cutting a plasmid with two different enzymes prevents the plasmid from reforming a circle because the circle can't reform if the two different sides don't fit together. If it was only one enzyme the two sides will just fit together eventually.
In this step, you are asked to set up a tube without
the restriction enzymes, BamHI and HindIII. What is the purpose of this
step, and why is it important?
So we can keep the lab controlled and it is important because we can base our answers off it.
Why might the enzymes work best at 37°C? Why
should the enzymes then be placed in the freezer? (Hint: The human body
temperature is 37°C.)
It can grow the rate as if it was in your body and we would put the enzymes in the freezer so they wouldn't grow any more.
Lab 4a
The DNA is not visible as it moves through the gel. The loading dye contains the three dyes that you separated in Laboratory 1.2. Why is it useful to use the loading dye in this lab? So we could see the DNA move from one side to the other on the gel.
Lab 5a
Make RFP from jelly fish in bacteria
Learn about steps of genetic engineering
Materials and Procedure
2a- materials and procedure can be found in Amgen manual part 2a
4a- materials and procedure can be found in Amgen manual part 4a
5a- materials and procedure can be found in Amgen manual part 5a
6a- materials and procedure can be found in Amgen manual part 6a
Experimental Overview
Part 2a: Use restriction digest to verify plasmid. We cut the plasmid with Hindlll and BamH1, to have the ability to get a portion of the bacteria gene.
Part 4a: Verification of plasmid digest by electrophoresis. We used gel electrophoresis in which a plasmid is ran along a DNA ladder. This ladder has bands that go hand-in-hand with other spaces or lane to determine the molecules stature. We know the size of the RFP gene so we knew how well the digest had ran. The digest ran perfectly.
Part 5a: Transformation of bacteria with recombinant plasmid. We cut plasmids with enzymes specifically restriction enzymes and to put together the gene of interest we used a substance called ligase.
Part 6a: We incubated the RFP gene or kept it warm and we separated the RFP from everything else. Next we used hydrophilic beads to collect the RFP and finally we transported the RFP and sort of made it mad by striking it against a pipet rack.
Data
2a- Why is it important that the same enzyme or
enzymes be used to cut both the plasmid and the insulin gene from the
human DNA?
If we cut the plasmid and the insulin gene from the DNA so the ends of them connect and fit together like puzzle pieces.
Why does using two different enzymes to cut the
plasmid prevent the plasmid from reforming a circle without the
inserted gene?
Cutting a plasmid with two different enzymes prevents the plasmid from reforming a circle because the circle can't reform if the two different sides don't fit together. If it was only one enzyme the two sides will just fit together eventually.
In this step, you are asked to set up a tube without
the restriction enzymes, BamHI and HindIII. What is the purpose of this
step, and why is it important?
So we can keep the lab controlled and it is important because we can base our answers off it.
Why might the enzymes work best at 37°C? Why
should the enzymes then be placed in the freezer? (Hint: The human body
temperature is 37°C.)
It can grow the rate as if it was in your body and we would put the enzymes in the freezer so they wouldn't grow any more.
- List in words or indicate in a drawing the important features of a plasmid vector that are required to clone a gene. Explain the purpose of each feature. Important features of a plasmid vector include the sequence that start DNA replication, which can allow the plasmid to replicate its self. Also the gene encodes a protein for an antibodies resistance. This allows the plasmid to identify bacteria. Finally a promoter sequence which starts the transcription of an inserted, specific gene
- What role do restriction enzymes have in nature? They protect the bacteria from infections such as viruses.
- Using your understanding of evolution, why would bacteria retain a gene that gives them resistance to antibiotics? How is the existence of bacteria with antibiotic resistance affecting medicine today? The bacteria with the resistance to antibodies will reproduce much more because of there advantage over the bacteria without the antibiotic resistance gene. The bacteria are being mutated and now they can't be killed by some antibiotics.
- Bacteria, sea anemones, and humans seem, on the surface, to be very different organisms. Explain how a gene from humans or a sea anemone can be expressed in bacteria to make a product never before made in bacteria. A gene is matched with a promoter or bacterial promoter on a plasmid and the plasmid is received by the bacteria and this bacteria can make proteins.
- Due to a mishap in the lab, bacteria carrying a plasmid with an ampicillin- resistant gene and bacteria carrying a plasmid with a gene that provides resistance to another antibiotic (kanamycin) were accidentally mixed together. Design an experiment that will allow you to sort out the two kinds of bacteria. (Hint: Make sure that you do not kill off one of the kinds of bacteria you are trying to sort out!) Make one batch one with kanamycin and one with ampicilin and repeat all the steps we did and this can be a way to separate bacteria. Cut them each with a different insulin and plasmid gene.
Lab 4a
The DNA is not visible as it moves through the gel. The loading dye contains the three dyes that you separated in Laboratory 1.2. Why is it useful to use the loading dye in this lab? So we could see the DNA move from one side to the other on the gel.
- Why is it important to verify that you have the correct recombinant plasmid? If you don't have the correct recombinant plasmid the damaged bacteria will go unnoticed because it might not be visible, so you might have to redo the entire lab.
- How did your actual gel results compare to your gel predictions? my gel results were the same as my predictions because they have the same number of bands in each lane as I had predicted.
- Do you see any bands that are not expected? What could explain the origin of these unexpected bands? There is a couple of bands unexpected and this could be a result of either human error or we damaged the gel with the DNA.
- Does the gel show that you are using the correct recombinant plasmid? Describe the evidence you used to make this assessment. Yes because we got the correct results and the R+ lanes had two fragments, which was desired.
- In the R– lane, do you see evidence of multiple configurations of plasmids? Explain your answer. Yes I saw multiple different bands showing that the plasmid was digested.
- In the R+ lane, do you see evidence of complete digestion? Explain your answer. Yes once again there were multiple bands showing the plasmid was digested into multiple fragments.
- In which lane would you expect to find the rfp gene and the ampR gene in the gel photograph? Are you able to locate these two genes? Explain your answer. The rfp gene would be found in the R+ lane and the ampR gene in R- lane.We can locate these in both of these lanes.
- Compare the lanes that have linear fragments with the lanes that have plasmids. Is there a difference in the shape of the bands between these two DNA forms? The lanes with plasmids are supercoiled and both migrate in some way. Yes the fragments have elongated edges while the plasmids are supercoiled.
Lab 5a
- How is the P+ bacteria culture treated differently from the P– bacteria culture? (A culture is an isolated population of cells.) What is the purpose of the P– bacteria culture? The P+ was given a plasmid while P- wasn't. The P- bacteria culture is meant to become a plasmid controller.
- Why do the cells need time to recover after the heat shock? They need to recuperate and express its antibiotic resistance and they need to rest.
- Why are the cells incubated at 37°C? To create an environment much like the human body to have the ability to grow like cell division
- You will use aseptic technique in this lab. Why is this important? So we can control the experiment and keep the substance from being contaminated.
5a Questions- Look at the results of your transformation. Do your actual results match your predicted results? If not, what differences do you see, and what are some explanations for these differences? Our predictions were correct but we might have accidentally cross contaminated, but doubtful.
- How many red colonies were present on your LB/amp/ara plate? I don't think we had any.
- Why did the red colonies only appear on the LB/amp/ara plate and not the LB/amp plate? They didn't show up.
- Recombinant plasmids are engineered so that they can replicate in the cell independently of the chromosome replication. Why is it important to have multiple copies of a recombinant plasmid within a cell? Most of the time these are damaged, but if you have a lot we have a better chance of getting the correct answer.
- How is the information encoded in the rfp gene expressed as a trait? Be sure to use what you have previously learned about gene expression and the relationship between DNA, RNA, protein, and traits. RFP glows in the gel/ protein.
- Why is it possible for bacteria to make a human protein, such as insulin, or a sea anemone protein, such as the red fluorescent dye?They have similar compositions as us in terms of structure or organelles.
Lab 6
How can you determine where the red fluorescent protein is in each separation step? Because it shows up light pink to where we can see it.
What color is the supernatant? The pellet? What are the contents of each? The supernatant is light pink and the pellets were pink. The supernatants consisted of EC and the pellets consisted of RFP.
Three buffers you will use in this lab are the binding buffer (BB), the wash buffer (WB), and the elution buffer (EB). What is the function of each? Binding buffer=Binds proteins in EC Wash buffer= Wash unbound protein Elution buffer= knock RFP off column- Why is a protein’s conformation important for carrying out its function? In binding spaces proteins attach to other molecules and this allows it to carry out its function.
- What properties of the amino acids in a protein relate to protein folding? The sequence of the amino acids relates to folding.
- Does the eluate containing your red fluorescent protein appear less bright
or brighter than it did in the cell lysate following centrifugation? If there is a noticeable difference in the intensity of the red color, what might account for that? The Elute is brighter than the lysate. The RFP will account for that. - What characteristic of red fluorescent protein is used as the basis for separation by column chromatography? Hydrophobic amino acids
- How might the column chromatography procedure be adjusted or modified to increase the purity of the red fluorescent protein sample? Use more of the original buffer.
Analysis/Conclusion
Our red conies didn't show up so our experiment went wrong at some point. Maybe we cross-contaminated one of the buffers but I don't think so. Our R+ and R- met our predictions so we didn't go wrong there, but I would further like to research what we did wrong.
Reflection
I personally liked our group because our teacher picked them for us which allowed for new experiences with different people. I would have been more careful pipetting because thats where I think we went wrong. I also liked the packet even though it did have a lot of information I learned a lot. The only problem was it might have needed a little more explaining but I really liked how different this lab was compared to everything else that we have done.- How is the P+ bacteria culture treated differently from the P– bacteria culture? (A culture is an isolated population of cells.) What is the purpose of the P– bacteria culture? The P+ was given a plasmid while P- wasn't. The P- bacteria culture is meant to become a plasmid controller.
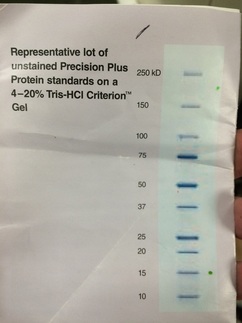
Final Gel Analysis
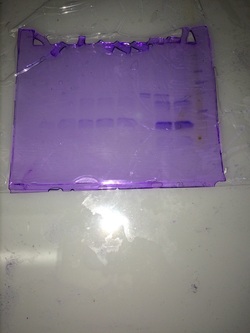
There are many lanes in this particular gel, but occupied lane 7. Compared to the Standards ours was just about average but it is harder to identify with this particular picture. Below you can see the standards for protein and ours was about 15-20 kD. Our stain was very pure, but not concentrated at all. We can find out the purity by seeing how many bands it has. We can see how large the band is by referencing the ladder or chart. We can see the concentration by how dark these bands or band is.